You can use boolean logic (e.g. AND/OR/NOT) for complex search queries. For more help and examples, see the search documentation.
Search by package name:
my-package (implicit)
name:my-package (explicit)
Search by package filename:
filename:my-package.ext
Search by package tag:
tag:latest
Search by package version:
version:1.0.0
prerelease:true (prereleases)
prerelease:false (no prereleases)
Search by package architecture:
architecture:x86_64
Search by package distribution:
distribution:el
Search by package license:
license:MIT
Search by package format:
format:deb
Search by package status:
status:in_progress
Search by package file checksum:
checksum:5afba
Search by package security status:
severity:critical
Search by package vulnerabilities:
vulnerabilities:>1
vulnerabilities:<1000
Search by # of package downloads:
downloads:>8
downloads:<100
Search by package type:
type:binary
type:source
Search by package size (bytes):
size:>50000
size:<10000
Search by dependency name/version:
dependency:log4j
dependency:log4j=1.0.0
dependency:log4j>1.0.0
Search by uploaded date:
uploaded:>"1 day ago"
uploaded:<"August 14, 2022 EST"
Search by entitlement token (identifier):
entitlement:3lKPVJPosCsY
Search by policy violation:
policy_violated:true
deny_policy_violated:true
license_policy_violated:true
vulnerability_policy_violated:true
Search by repository:
repository:repo-name
Search by last download date:
last_downloaded:<"30 days ago"
last_downloaded:>"August 14, 2022 EST"
Search queries for all Debian-specific (and related) package types
Search by component:
deb_component:unstable
Search queries for all Maven-specific (and related) package types
Search by group ID:
maven_group_id:org.apache
Search queries for all Docker-specific (and related) package types
Search by image digest:
docker_image_digest:sha256:7c5..6d4
(full hashref only)
Search by layer digest:
docker_layer_digest:sha256:4c4..ae4
(full hashref only)
Search queries for all Generic-specific package types
Search by file path:
generic_filepath:path/to/file.txt
Search by directory:
generic_directory:path/to
Field type modifiers (depending on the type, you can influence behaviour)
For all queries, you can use:
~foo for negation
For string queries, you can use:
^foo to anchor to start of term
foo$ to anchor to end of term
foo*bar for fuzzy matching
For number/date or version queries, you can use:
>foo for values greater than
>=foo for values greater / equal
<foo for values less than
<=foo for values less / equal
Need a secure and centralised artifact repository to deliver Alpine,
Cargo,
CocoaPods,
Composer,
Conan,
Conda,
CRAN,
Dart,
Debian,
Docker,
Generic,
Go,
Helm,
Hex,
HuggingFace,
LuaRocks,
Maven,
npm,
NuGet,
P2,
Python,
RedHat,
Ruby,
Swift,
Terraform,
Vagrant,
VSX,
Raw & More packages?
Cloudsmith is the new standard in Package / Artifact Management and Software Distribution.
With support for all major package formats, you can trust us to manage your software supply chain.
 brainreg
1.0.5
brainreg
1.0.5
One-liner (summary)
Description
[](https://pypi.org/project/brainreg) [](https://pypi.org/project/brainreg) [](https://pypi.org/project/brainreg) [](https://github.com/brainglobe/brainreg) [](https://github.com/brainglobe/brainreg/actions) [](https://codecov.io/gh/brainglobe/brainreg) [](https://github.com/python/black)
# brainreg
brainreg is an update to [amap](https://github.com/SainsburyWellcomeCentre/amap_python) (which is itself a port of the [original Java software](https://www.nature.com/articles/ncomms11879)) to include multiple registration backends, and to support the many atlases provided by [bg-atlasapi](https://github.com/brainglobe/bg-atlasapi). It also comes with an optional [napari plugin](https://github.com/brainglobe/brainreg-napari) if you'd rather use brainreg with through graphical interface.
Documentation for both the command-line tool and graphical interface can be found [here](https://brainglobe.info/documentation/brainreg/index.html). If you have any issues, please get in touch [on the forum](https://forum.image.sc/tag/brainglobe), [on Zulip](https://brainglobe.zulipchat.com/), or by [raising an issue](https://github.com/brainglobe/brainreg/issues).
For segmentation of bulk structures in 3D space (e.g. injection sites, Neuropixels probes), please see [brainglobe-segmentation](https://github.com/brainglobe/brainglobe-segmentation).
## Details
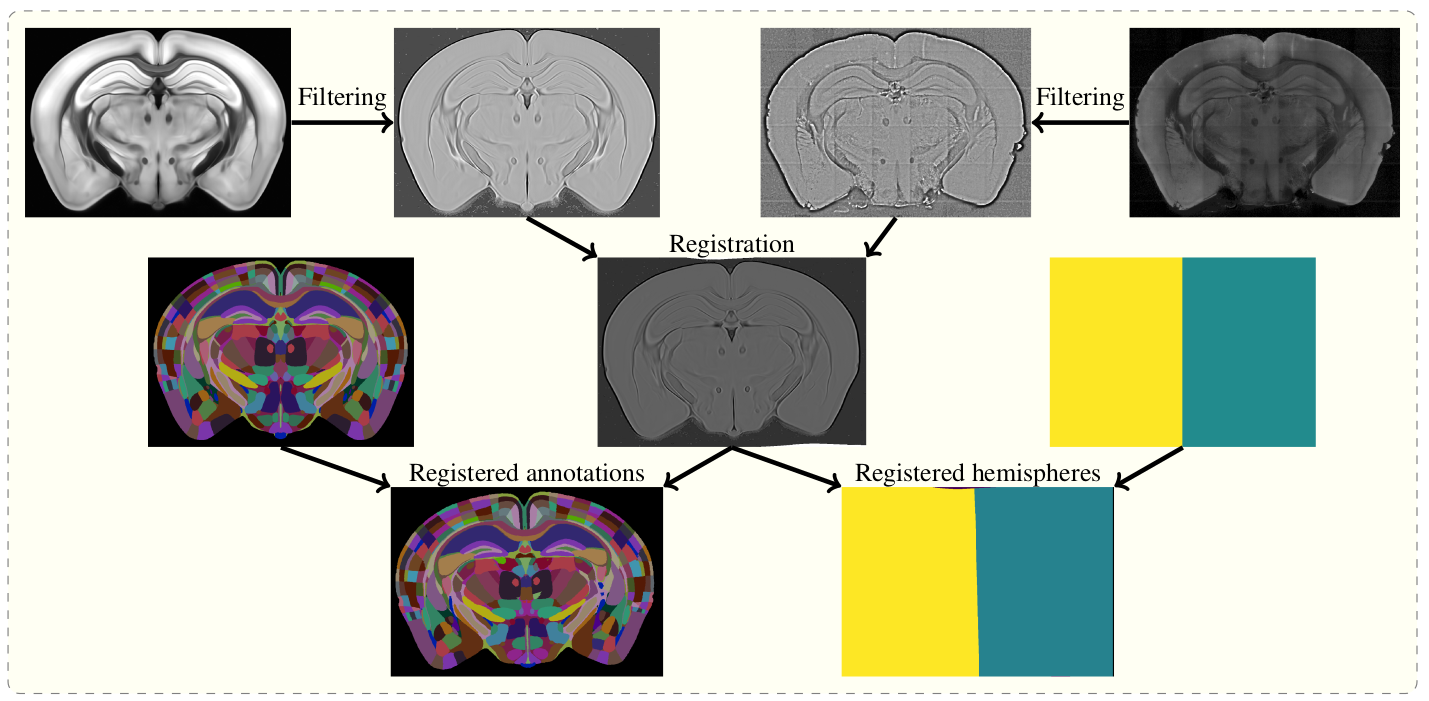
The aim of brainreg is to register the template brain (e.g. from the [Allen Reference Atlas](https://mouse.brain-map.org/static/atlas)) to the sample image. Once this is complete, any other image in the template space can be aligned with the sample (such as region annotations, for segmentation of the sample image). The template to sample transformation can also be inverted, allowing sample images to be aligned in a common coordinate space.
To do this, the template and sample images are filtered, and then registered in a three step process (reorientation, affine registration, and freeform registration). The resulting transform from template to standard space is then applied to the atlas.
Full details of the process are in the [original aMAP paper](https://www.nature.com/articles/ncomms11879).
## Installation
To install both the command line tool and the napari plugin, run
`bash pip install brainreg[napari] `
in your desired Python environment. To only install the command line tool with no GUI (e.g. to run brainreg on an HPC cluster), just run:
`bash pip install brainreg `
### Installing on macOS
If you are using macOS, please run
`bash conda install -c conda-forge niftyreg `
in your environment before installing, to ensure all dependencies are installed.
## Command line usage
### Basic usage
`bash brainreg /path/to/raw/data /path/to/output/directory -v 5 2 2 --orientation psl `
Full command-line arguments are available with brainreg -h, but please [get in touch](mailto:code@adamltyson.com?subject=brainreg) if you have any questions.
### Mandatory arguments
- Path to the directory of the images. This can also be a text file pointing to the files.
- Output directory for all intermediate and final results.
- You must also specify the voxel sizes with the -v flag, see [specifying voxel size](https://brainglobe.info/documentation/general/image-definition.html#voxel-sizes) for details.
### Atlas
By default, brainreg will use the 25um version of the [Allen Mouse Brain Atlas](https://mouse.brain-map.org/). To use another atlas (e.g. for another species, or another resolution), you must use the --atlas flag, followed by the string describing the atlas, e.g.:
`bash --atlas allen_mouse_50um `
To find out which atlases are available, once brainreg is installed, please run brainglobe list. The name of the resulting atlases is the string to pass with the --atlas flag.
### Input data orientation
If your data does not match the BrainGlobe default orientation (the origin voxel is the most anterior, superior, left-most voxel), then you must specify the orientation by using the --orientation flag. What follows must be a string in the [brainglobe-space](https://github.com/brainglobe/brainglobe-space) "initials" form, to describe the origin voxel.
If the origin of your data (first, top left voxel) is the most anterior, superior, left part of the brain, then the orientation string would be "asl" (anterior, superior, left), and you would use:
`bash --orientation asl `
### Registration options
To change how the actual registration performs, see [registration parameters](https://brainglobe.info/documentation/brainreg/user-guide/parameters.html)
### Additional options
- -a or --additional Paths to N additional channels to downsample to the same coordinate space.
- --sort-input-file If set to true, the input text file will be sorted using natural sorting. This means that the file paths will be sorted as would be expected by a human and not purely alphabetically.
- --brain_geometry Can be one of full (default) for full brain registration, hemisphere_l for left hemisphere data-set and hemisphere_r for right hemisphere data-set.
### Misc options
- --n-free-cpus The number of CPU cores on the machine to leave unused by the program to spare resources.
- --debug Debug mode. Will increase verbosity of logging and save all intermediate files for diagnosis of software issues.
- --save-original-orientation Option to save the registered atlas with the same orientation as the input data.
## Visualising results
If you have installed the optional [napari](https://github.com/napari/napari) plugin, you can use napari to view your data. The plugin automatically fetches the [brainglobe-napari-io](https://github.com/brainglobe/brainglobe-napari-io) which provides this functionality. If you have installed only the command-line tool you can still manually install [brainglobe-napari-io](https://github.com/brainglobe/brainglobe-napari-io) and follow the steps below.
### Sample space
Open napari and drag your brainreg output directory (the one with the log file) onto the napari window.
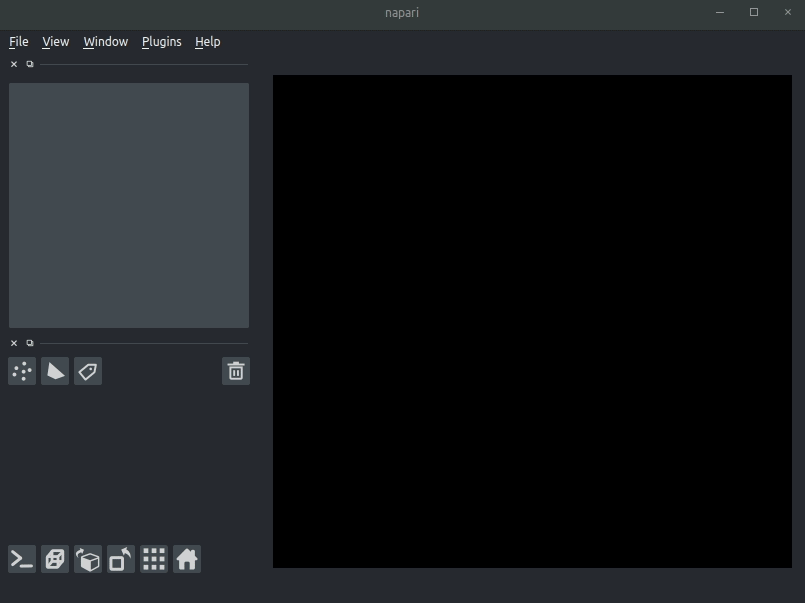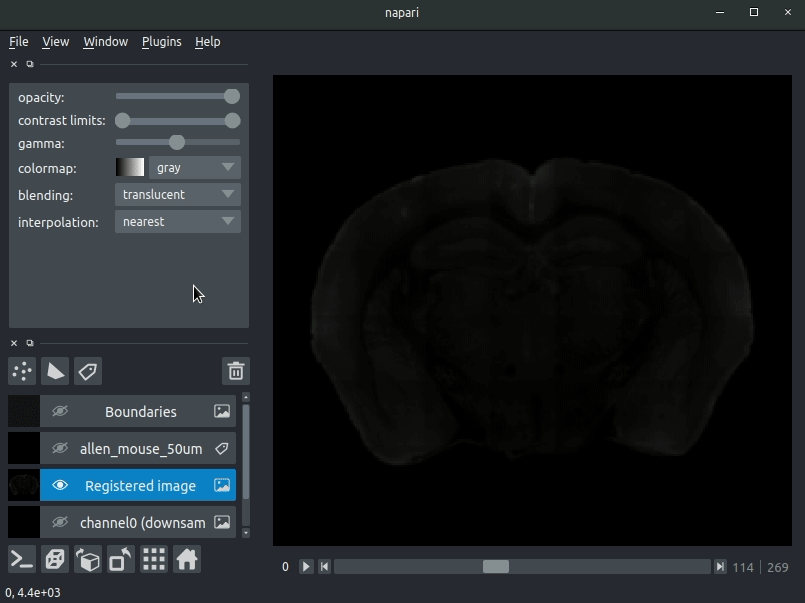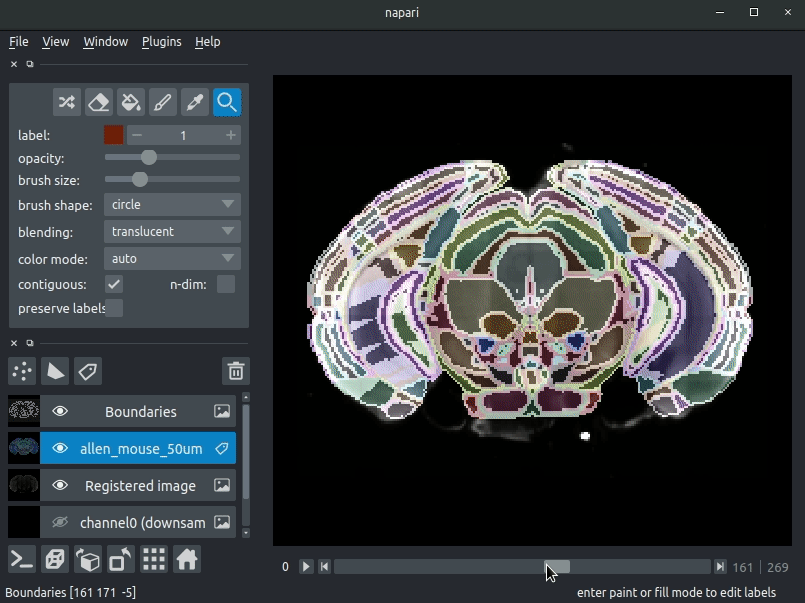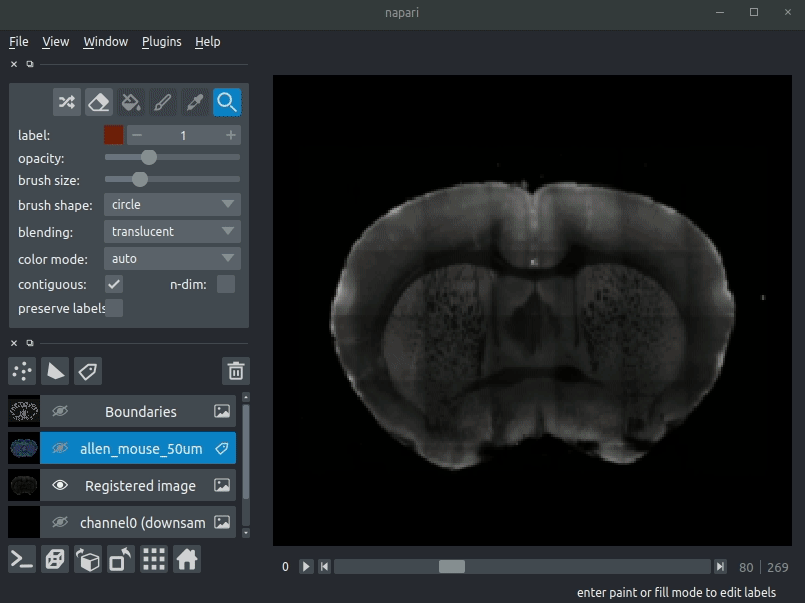
Various images should then open, including:
- Registered image - the image used for registration, downsampled to atlas resolution
- atlas_name - e.g. allen_mouse_25um the atlas labels, warped to your sample brain
- Boundaries - the boundaries of the atlas regions
If you downsampled additional channels, these will also be loaded. Most of these images will not be visible by default - click the little eye icon to toggle visibility.
Note: If you use a high resolution atlas (such as allen_mouse_10um), then the files can take a little while to load.
## Contributing
Contributions to brainreg are more than welcome. Please see the [developers guide](https://brainglobe.info/developers/index.html).
## Citing brainreg
If you find brainreg useful, and use it in your research, please let us know and also cite the paper:
> Tyson, A. L., Vélez-Fort, M., Rousseau, C. V., Cossell, L., Tsitoura, C., Lenzi, S. C., Obenhaus, H. A., Claudi, F., Branco, T., Margrie, T. W. (2022). Accurate determination of marker location within whole-brain microscopy images. Scientific Reports, 12, 867 [doi.org/10.1038/s41598-021-04676-9](https://doi.org/10.1038/s41598-021-04676-9)
Please also cite aMAP (the original pipeline from which this software is based):
>Niedworok, C.J., Brown, A.P.Y., Jorge Cardoso, M., Osten, P., Ourselin, S., Modat, M. and Margrie, T.W., (2016). AMAP is a validated pipeline for registration and segmentation of high-resolution mouse brain data. Nature Communications. 7, 1–9. <https://doi.org/10.1038/ncomms11879>
Lastly, if you can, please cite the BrainGlobe Atlas API that provided the atlas:
>Claudi, F., Petrucco, L., Tyson, A. L., Branco, T., Margrie, T. W. and Portugues, R. (2020). BrainGlobe Atlas API: a common interface for neuroanatomical atlases. Journal of Open Source Software, 5(54), 2668, <https://doi.org/10.21105/joss.02668>
Finally, don't forget to cite the developers of the atlas that you used (e.g. the Allen Brain Atlas)!
| Status | Completed |
|---|---|
| Checksum (MD5) | 179389fae09dfbc750952cc6092ac8b7 |
| Checksum (SHA-1) | 5b98a3bb99c1c410537b045014e4e4b36459b85a |
| Checksum (SHA-256) | 4556b912c555a871e5fef2a3c045fe0a6ba412d8049f53226b86c0816ee34a5d |
| Checksum (SHA-512) | c886877e5dd860d638e700e8fa0e24f6d22a4b09e9bf2f243d637b7220ccc1e7f5… |
| GPG Signature | |
| GPG Fingerprint | 6811684bac0b8895434e97bdd4391b8fb999e537 |
| Storage Region | Dublin, Ireland |
| Type | Binary (contains binaries and binary artifacts) |
| Uploaded At | 4 months, 3 weeks ago |
| Uploaded By |
|
| Slug Id | brainreg-105-py3-none-anywhl-bvyt |
| Unique Id | qd5FigZncjk2mka3 |
| Version (Raw) | 1.0.5 |
| Version (Parsed) |
|
| extended metadata | |
| Author | Adam Tyson, Charly Rousseau, Stephen Lenzi <code@adamltyson.com> |
| Classifiers | Development Status :: 3 - Alpha | Framework :: napari | Intended Audience :: Developers | Intended Audience :: Science/Research | Operating System :: OS Independent | Programming Language :: Python | Programming Language :: Python :: 3.10 | Programming Language :: Python :: 3.11 | Programming Language :: Python :: 3.9 |
| Metadata Version | 2.1 |
| Project Urls | Bug Tracker, https://github.com/brainglobe/brainreg/issues | Documentation, https://docs.brainglobe.info/brainreg | Homepage, https://brainglobe.info | Source Code, https://github.com/brainglobe/brainreg | Twitter, https://twitter.com/brain_globe | User support, https://forum.image.sc/tag/brainglobe |
| Py Filetype | bdist_wheel |
| Py Version | py3 |
| Requires Dist | bg-atlasapi | black ; extra == 'dev' | brainglobe-napari-io >=0.3.2 ; extra == 'napari' | brainglobe-segmentation >=1.0.0 ; extra == 'napari' | brainglobe-space >=1.0.0 | brainglobe-utils >=0.4.0 | check-manifest ; extra == 'dev' | fancylog | gitpython ; extra == 'dev' | magicgui ; extra == 'napari' | napari-plugin-engine >=0.1.4 ; extra == 'napari' | napari[pyqt5] ; extra == 'dev' | napari[pyqt5] ; extra == 'napari' | numpy | pooch >1 ; extra == 'napari' | pre-commit ; extra == 'dev' | pytest ; extra == 'dev' | pytest-cov ; extra == 'dev' | pytest-qt ; extra == 'dev' | qtpy ; extra == 'napari' | scikit-image | setuptools-scm ; extra == 'dev' | tox ; extra == 'dev' |
| Requires Python | >=3.9 |
| pkg | brainreg-1.0.5-py3-none-any.whl |
6
12.8 MB |
md5 | sha1 | sha256 | sha512 |
This package has 71 files/directories.
| Newer |
|
brainreg |
10 |
|
||
| Newer |
|
brainreg |
8 |
|
||
| Newer |
|
brainreg |
8 |
|
||
| Newer |
|
brainreg |
8 |
|
||
| Newer |
|
brainreg |
9 |
|
||
| Newer |
|
brainreg |
8 |
|
||
| Newer |
|
brainreg |
8 |
|
||
| Newer |
|
brainreg |
10 |
|
||
| Newer |
|
brainreg |
9 |
|
||
|
|
brainreg |
6 |
|
|||
| Older |
|
brainreg |
7 |
|
||
| Older |
|
brainreg |
8 |
|
||
| Older |
|
brainreg |
9 |
|
||
| Older |
|
brainreg |
11 |
|
||
| Older |
|
brainreg |
9 |
|
||
| Older |
|
brainreg |
9 |
|
||
| Older |
|
brainreg |
5 |
|
||
| Older |
|
brainreg |
6 |
|
||
| Older |
|
brainreg |
7 |
|
||
| Older |
|
brainreg |
8 |
|
Last scanned
4 months, 3 weeks ago
Scan result
Clean
Vulnerability count
0
Max. severity
UnknownPackage statistics are no longer available on cloudsmith.io. Please visit our new web app to access this feature.
You can embed a badge in another website that shows this or the latest version of this package.
To embed the badge for this specific package version, use the following:
[](https://cloudsmith.io/~demo-docs/repos/awesome-repo/packages/detail/python/brainreg/1.0.5/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/)|This version of 'brainreg' @ Cloudsmith|
.. |This version of 'brainreg' @ Cloudsmith| image:: https://api.cloudsmith.com/v1/badges/version/demo-docs/awesome-repo/python/brainreg/1.0.5/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/?render=true
:target: https://cloudsmith.io/~demo-docs/repos/awesome-repo/packages/detail/python/brainreg/1.0.5/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/image::https://api.cloudsmith.com/v1/badges/version/demo-docs/awesome-repo/python/brainreg/1.0.5/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/?render=true[link="https://cloudsmith.io/~demo-docs/repos/awesome-repo/packages/detail/python/brainreg/1.0.5/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/",title="This version of 'brainreg' @ Cloudsmith"]<a href="https://cloudsmith.io/~demo-docs/repos/awesome-repo/packages/detail/python/brainreg/1.0.5/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/"><img src="https://api.cloudsmith.com/v1/badges/version/demo-docs/awesome-repo/python/brainreg/1.0.5/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/?render=true" alt="This version of 'brainreg' @ Cloudsmith" /></a>rendered as:
To embed the badge for the latest package version, use the following:
[](https://cloudsmith.io/~demo-docs/repos/awesome-repo/packages/detail/python/brainreg/latest/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/)|Latest version of 'brainreg' @ Cloudsmith|
.. |Latest version of 'brainreg' @ Cloudsmith| image:: https://api.cloudsmith.com/v1/badges/version/demo-docs/awesome-repo/python/brainreg/latest/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/?render=true&show_latest=true
:target: https://cloudsmith.io/~demo-docs/repos/awesome-repo/packages/detail/python/brainreg/latest/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/image::https://api.cloudsmith.com/v1/badges/version/demo-docs/awesome-repo/python/brainreg/latest/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/?render=true&show_latest=true[link="https://cloudsmith.io/~demo-docs/repos/awesome-repo/packages/detail/python/brainreg/latest/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/",title="Latest version of 'brainreg' @ Cloudsmith"]<a href="https://cloudsmith.io/~demo-docs/repos/awesome-repo/packages/detail/python/brainreg/latest/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/"><img src="https://api.cloudsmith.com/v1/badges/version/demo-docs/awesome-repo/python/brainreg/latest/a=noarch;xf=bdist_wheel;xn=brainreg;xv=py3/?render=true&show_latest=true" alt="Latest version of 'brainreg' @ Cloudsmith" /></a>rendered as:
These instructions assume you have setup the repository first (or read it).
To install/use brainreg @ version 1.0.5 ...
pip install 'brainreg==1.0.5'You can also install the latest version of this package:
pip install --upgrade 'brainreg'If necessary, you can specify the repository directly:
pip install \
--index-url=https://dl.cloudsmith.io/public/demo-docs/awesome-repo/python/simple/ \
brainreg==1.0.5If you've got a project requirements.txt file, you can specify this as a dependency:
--index-url=https://dl.cloudsmith.io/public/demo-docs/awesome-repo/python/simple/
brainreg==1.0.5In addition, you can use this repository as an extra index url. However, please read our documentation on this parameter before using it. For example in a requirements.txt file:
--extra-index-url=https://dl.cloudsmith.io/public/demo-docs/awesome-repo/python/simple/
brainreg==1.0.5